Introduction:
|
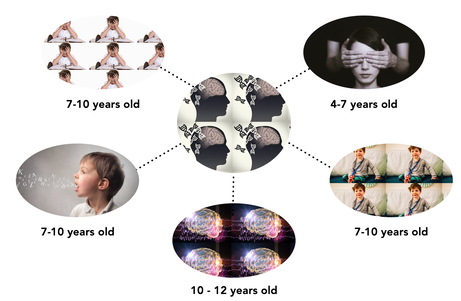
Batten Disease is a rare neurodegenerative disorder affecting 4 to 6 out of 100,000 children in the United States. A child born with Batten Disease has "normal" development until the age of four when the symptoms of the disease begin to appear. From ages four to seven, a child starts to lose their vision. From seven to ten years old, a child with Batten Disease will lose their cognitive abilities (shortened attention span, more tired, etc.), have decreased motor skills (such as walking or refined use of their hands), and increased difficulty to enunciate words. It is common that during this time a child might be misdiagnosed with Attention Deficit Disorder (ADD), or other common learning disabilities. From ages ten through twelve, a child begins to have seizures with epileptic-like patterns. Typically, a child with Batten Disease does not survive to twenty years of age.
|
CLN3 is most commonly associated with the juvenile form of Batten Disease. A deletion mutation of exons seven and eight in this gene is recessively inherited from two heterozygous parents and cause a child to have the disease.
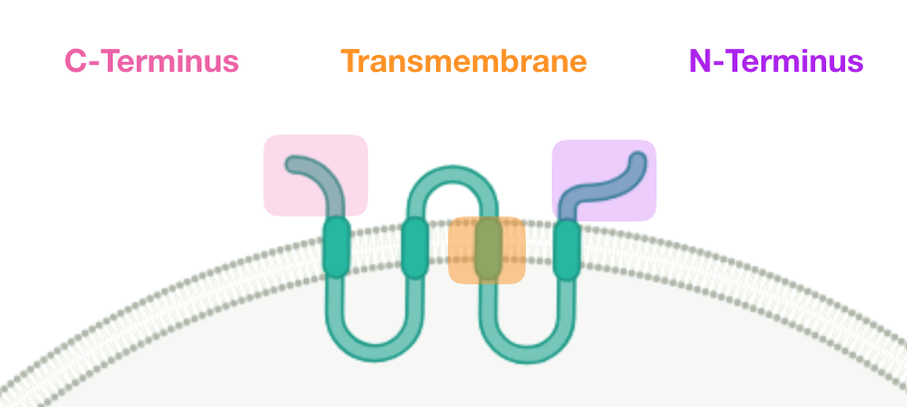
The CLN3 gene is a transmembrane protein found in the plasma membrane of the neuronal lysosome. Thus far, scientists have only conclusive information about the cellular component of this gene. The molecular and biological functions of the CLN3 protein are not yet entirely understood. Due to the location of the protein and the nature of organisms when the protein is mutated, it is believed that CLN3 is involved in cellular waste metabolism.
The CLN3 gene is a transmembrane protein found in the plasma membrane of the neuronal lysosome. Thus far, scientists have only conclusive information about the cellular component of this gene. The molecular and biological functions of the CLN3 protein are not yet entirely understood. Due to the location of the protein and the nature of organisms when the protein is mutated, it is believed that CLN3 is involved in cellular waste metabolism.
|
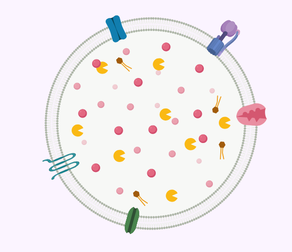
CLN3 is found in the plasma membrane of the neuronal lysosome. The function of this transmembrane protein is not yet known. The lysosome is known as the "trash can" of the cell, breaking down cellular waste from other organelles.
|
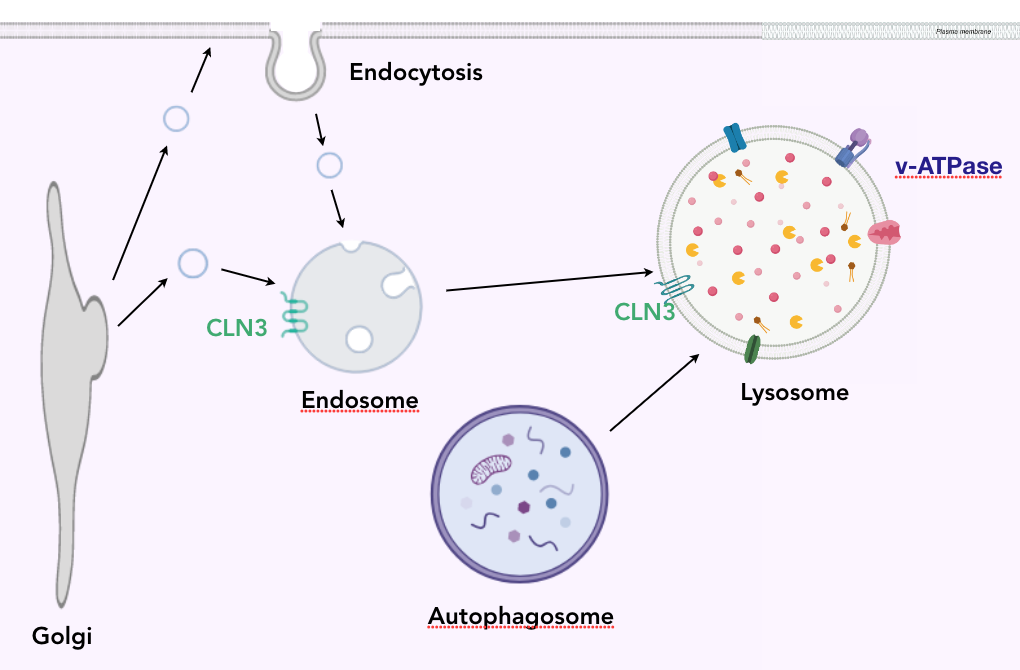
The CLN3 protein is synthesized in the endoplasmic reticulum, and moved to the Golgi. When expelled from the golgi in a vesicle, the CLN3 protein docks into the plasma membrane of the endosome. When the endosome and autophagosome merge with the lysosome, transmembrane proteins dock into the plasma membrane and enzymes reside in the lumen. The low pH of the lysosome creates a proton gradient which is useful for bringing waste into the organelle.
|
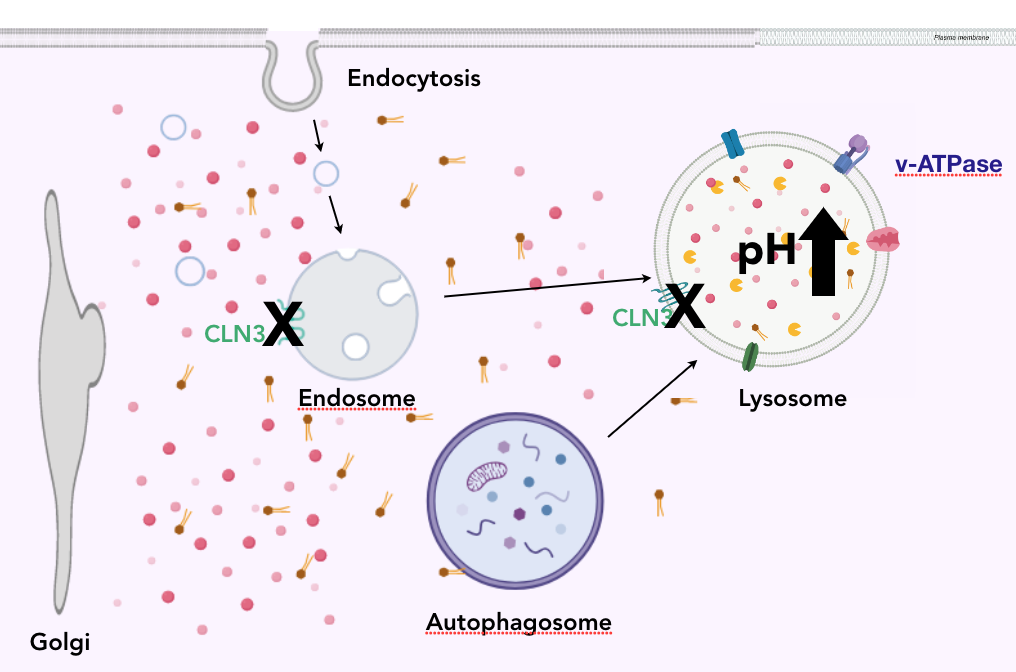
When CLN3 is mutated, the CLN3 protein is not functioning in the plasma membrane of the lysosome. When this occurs, the pH of the lysosome noticeably increases, and cellular waste begins to accumulate in the cytoplasm of the neuron. High concentrations of cellular waste in the cytoplasm hinders the functionality of the organelles, and eventually leads to cell death. The lowering of the pH is due to the inability of ATPase to move cationic amino acids into the lumen of the lysosome. The relationship between CLN3 and ATPase is being heavily studied.
|
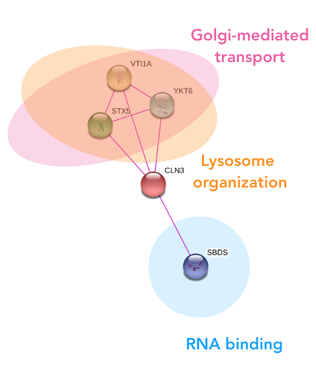
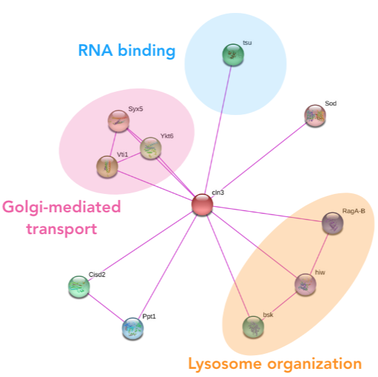
The protein interactions of CLN3 confirm the way in which CLN3 comes to reside in the plasma membrane of the neuronal lysosome, however, does not suggest a clear function of the protein. The direct interaction with the RNA binding protein was a very peculiar finding, and suggests that CLN3 could be involved in the translation or posttranslational modifications of other transport proteins in the plasma membrane of the neuronal lysosome. This interaction was conserved across humans and the model organism, suggesting the interaction with the RNA binding protein is key to its function.
Even with all the information presented above, the role of CLN3 in cellular waste metabolism leading to retinal degeneration is still unknown. This led to my primary goal of understanding the intricacies of CLN3 in cellular waste metabolism in neurons, and how mutations lead to retinal degeneration. I hypothesize that the CLN3 protein is involved with translation/post translational modifications of other transmembrane proteins that are responsible for cellular waste metabolism of neurons in the retina.
Model Organism
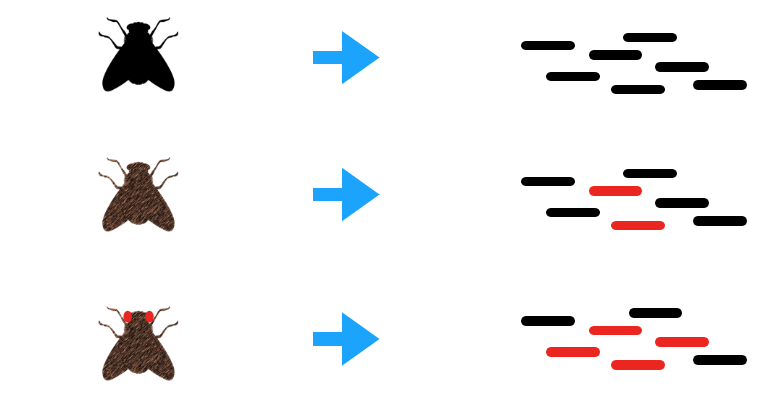
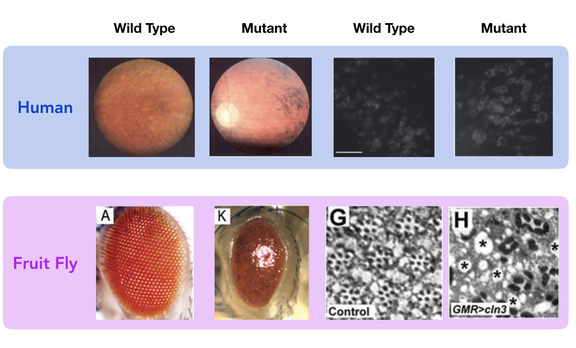 Figure 7. Comparison of CLN3 mutant and wild type phenotypes in Fruit flies and humans.
Figure 7. Comparison of CLN3 mutant and wild type phenotypes in Fruit flies and humans.
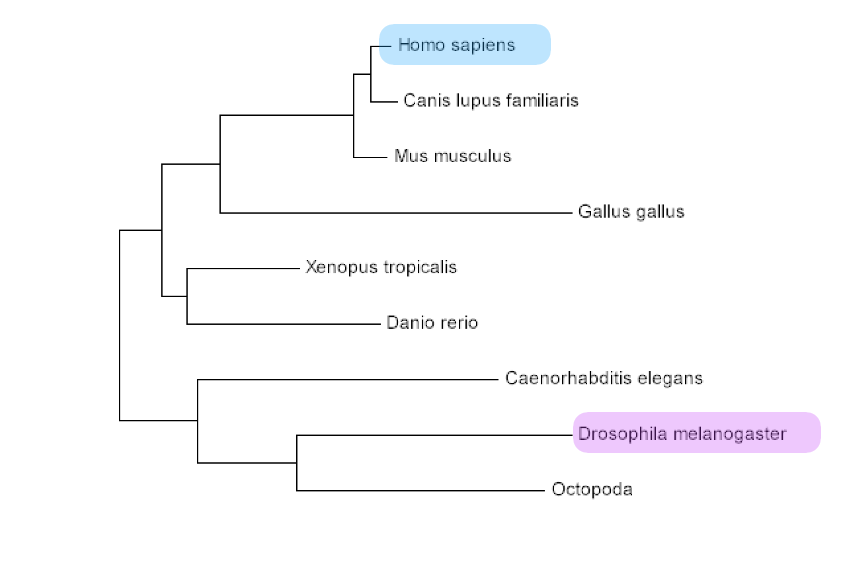
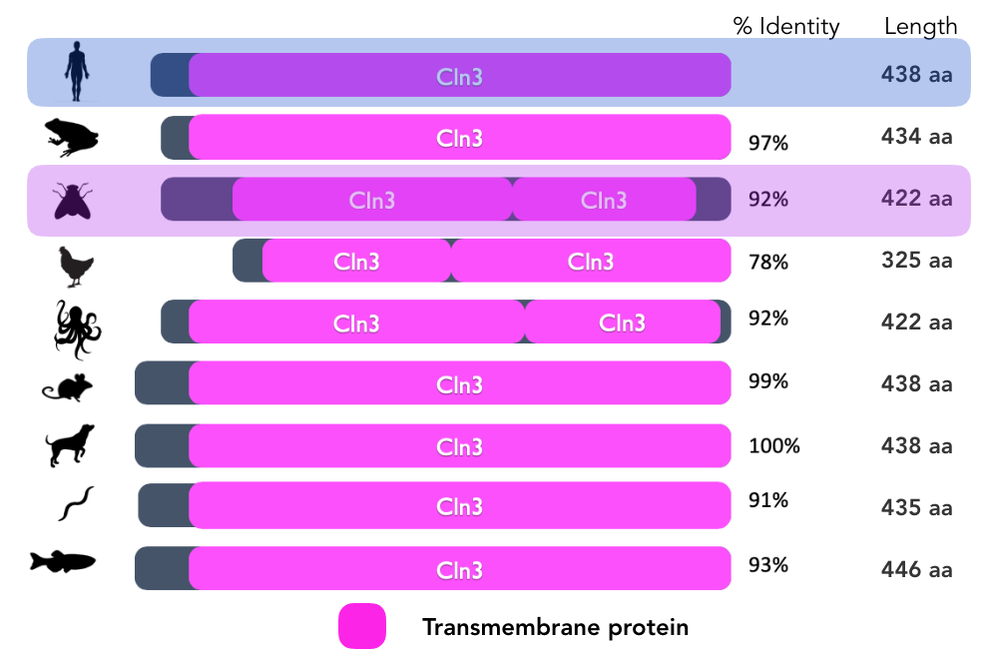
Drosophila melanogaster would be the best model organism while studying retinal degeneration in patients with Batten Disease. The phenotype of the mutant and wild type resemble that of humans in regards to neuronal waste buildup and retinal degeneration. Fruit flies are inexpensive and reproduce very quickly in great quantities. Also, the protein is highly conserved in this species and is not distantly related to the human form of the protein. Mice would not be an ideal model organism for this research, because mice lack a macular (center of the eye), meaning their anatomy could change the way that CLN3 impacts the function of the retina.
Aim One: Identify motifs that interact with RNA binding proteins in neuronal retina cells
|
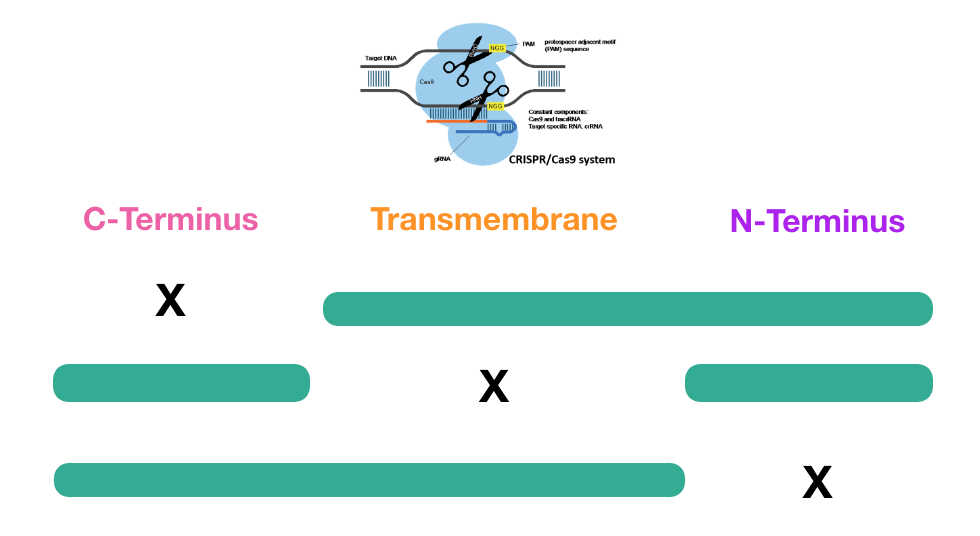
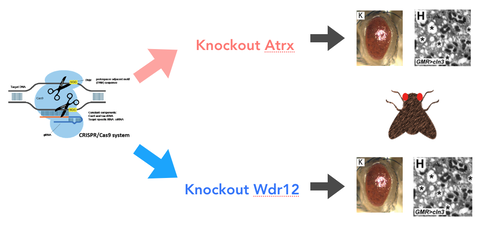
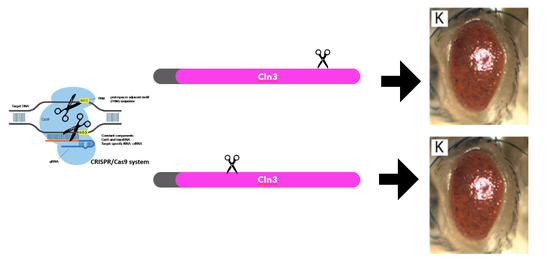
Although the findings in the protein interaction network suggest that CLN3 directly interacts with RNA binding proteins, it is still unknown how the CLN3 transmembrane protein conducts this interaction, as well as the result of the interaction. To solve at least one of these questions, I would like to conduct an experiment using CRISPR/Cas9 to create three separate deletions of CLN3 protein motifs in three separate trials. The first will be of the C-terminus motif, the second of the transmembrane region, and finally of the N-terminus motif. The result of this experiment would highlight which region of the CLN3 protein is responsible for interacting with the RNA binding proteins, such that the region responsible would have a mutant phenotype. If the other motifs do not interact with the RNA binding protein, then when deleted, the phenotype should resemble the wild type.
|
Because these experiments have not yet been conducted, I believe there could be three potential outcomes: wild type phenotype, partial mutant with cellular waste buildup and normal retina function, and finally a full mutant with both cellular waste buildup and mutant retina function. It is not yet known whether cellular waste buildup in the neurons is the direct cause of retinal degeneration, therefore the half mutant must be considered.
I hypothesize that the region responsible for interacting with RNA binding proteins is the C-terminus. Unlike the transmembrane region, the C-terminus and N-terminus are accessible to RNA binding proteins because they are not embedded within a plasma membrane, nor are enclosed within the lumen of the lysosome. There has also been significantly more research conducted on the C-terminus of the CLN3 protein than the N-terminus. The findings from some of the studies included the discovery of phosphorylation sites responsible for translation in the C-terminus. |
Aim Two: Characterize differences in gene transcription levels in mutated retina
|
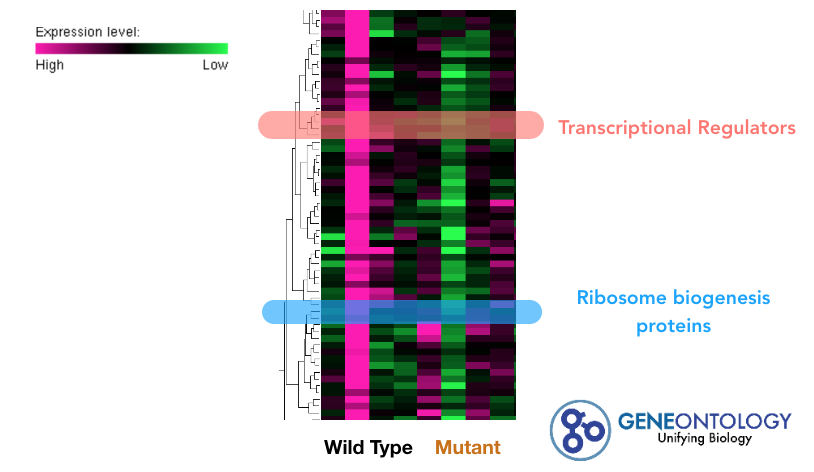
Next, I will conduct an experiment that will give insight to which genes are upregulated or downregulated when CLN3 is mutated. Because I am focusing on cellular waste metabolism in neurons leading to retinal degeneration, I will be retrieving samples from retinas of the eyes of the Drosophila. After retrieving samples from the wild type, half-mutant, and full mutant. The samples will be used for RNA sequencing (RNA-seq). These results will then be collected to form a heat map, which should directly show which genes are up or down regulated when CLN3 is mutated.
|
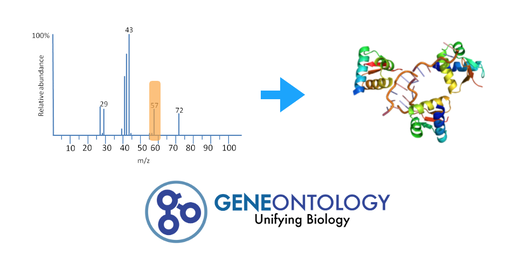
Using the information from the heat map, gene ontology terminology will be used to array the genes. Once a gene of interest is identified, CRISPR/Cas9 will be used to knockout the gene. This will determine if the gene will create the full-mutant phenotype. Such information would give insight to the pathway regulated by CLN3 as well as how cellular degeneration is connected to retinal degeneration.
I hypothesize that gene expression levels of RNA binding proteins responsible for translation of transmembrane proteins in the neuronal lysosome will be down-regulated in CLN3 mutants. |
Aim Three: Identify protein interactions of CLN3 involved in cellular waste metabolism in retina neurons.
|
In my final experiment, I would like to identify proteins that are specifically responsible for translation of transmembrane proteins important for trafficking cellular waste into the neuronal lysosome. Thus far, we know that CLN3 interacts with RNA binding proteins; however, we do not know the extent of the interactions, nor which proteins are translated from the RNA binding proteins that CLN3 interacts with. Some lysosomal transmembrane proteins are translated in the endoplasmic reticulum where CLN3 is also believed to originate. Therefore, CLN3 could be involved with the productivity of RNA binding proteins responsible for translating lysosomal transmembrane proteins. A mutation of CLN3 could prevent the proper formation of these proteins.I hypothesize that lysosomal transmembrane proteins responsible for transport of cellular waste will not be translated in CLN3 mutants, thus will cause a buildup of cellular waste in the neuron.
|
|
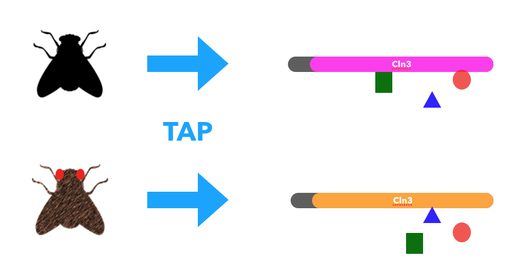
The experiment will begin with neuronal samples taken from the wild type, half-mutant, and full-mutant Drosophila eyes. These cells will be subjected to Tandem Affinity Purification (TAP), then Mass Spectrometry to determine the difference in protein-protein interactions. Gene Ontology will then be used on the RNA binding proteins to determine their protein-interaction networks. This information could highlight which RNA binding proteins interact with CLN3 as well as which proteins it translates. The RNA binding proteins that have significant changes between the wild type, and CLN3 full-mutant will be knocked out and assayed. The results will confirm which RNA binding proteins are specifically regulated by CLN3 and give clear pathology of how mutant CLN3 leads to the buildup of cellular waste in neuronal retina cells.
|
Future Directions
In my research, I found information that ATPase is involved in pumping cationic amino acids into the neuronal lysosome. This was found to be dysfunctional when CLN3 was mutated [5]. When ATPase is dysfunctional, the pH of the lysosomal lumen increases, destroying the proton gradient necessary to move cellular waste into the lysosome. The interaction between CLN3 and ATPase is still highly unknown. I would like to continue my research on CLN3, investigating the relationship between CLN3 and the translation of ATPase specifically.
Final Talk Final
| zanetakos_final_talk_draft_revised_5_09_19.key | |
| File Size: | 9118 kb |
| File Type: | key |
| zanetakos_final_talk_draft_4_25_19__1_.key | |
| File Size: | 9224 kb |
| File Type: | key |
Final Talk Drafts
| zanetakos_final_talk_draft_4_5_19.key | |
| File Size: | 6932 kb |
| File Type: | key |
| zanetakos_final_talk_draft_4_5_19.pptx | |
| File Size: | 5030 kb |
| File Type: | pptx |
References:
Mathavarajah, S., Mclaren, M. D., & Huber, R. J. (2018). Cln3 function is linked to osmoregulation in a Dictyostelium model of Batten disease. Biochimica Et Biophysica Acta (BBA) -Molecular Basis of Disease,1864(11), 3559-3573. doi:10.1016/j.bbadis.2018.08.013
Somogyi, A., Petcherski, A., Beckert, B., Huebecker, M., Priestman, D., Banning, A., . . . Tikkanen, R. (2018). Altered Expression of Ganglioside Metabolizing Enzymes Results in GM3 Ganglioside Accumulation in Cerebellar Cells of a Mouse Model of Juvenile Neuronal Ceroid Lipofuscinosis. International Journal of Molecular Sciences,19(2), 625. doi:10.3390/ijms19020625
Boswell-Casteel, R. C., & Hays, F. A. (2016). Equilibrative nucleoside transporters-A review. Nucleosides, nucleotides & nucleic acids, 36(1), 7-30.
Perland, E., Bagchi, S., Klaesson, A., & Fredriksson, R. (2017). Characteristics of 29 novel atypical solute carriers of major facilitator superfamily type: evolutionary conservation, predicted structure and neuronal co-expression. Open biology, 7(9), 170142.
Schultz, M. L., Tecedor, L., Lysenko, E., Ramachandran, S., Stein, C. S., & Davidson, B. L. (2018). Modulating membrane fluidity corrects Batten disease phenotypes in vitro and in vivo. Neurobiology of disease, 115, 182-193.
Images:
https://www.verywellhealth.com/myotonic-muscular-dystrophy-symptoms-4570998
https://www.ninds.nih.gov/Disorders/Patient-Caregiver-Education/Fact-Sheets/Batten-Disease-Fact-Sheet
https://www.researchgate.net/figure/RNA-seq-data-analysis-of-BMs-AMs-and-TMs-A-Heatmap-of-hierarchical-clustering_fig1_321441357
http://geneontology.org/
https://www.researchgate.net/figure/Effect-of-cisd2-Ppt1-and-cln3-on-external-eye-morphology-Light-microscopic-images-A-K_fig10_261408029
https://www.studyread.com/importance-of-lysosomes/
https://www.sciencedirect.com/science/article/pii/S0167488907000705
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4201705/
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5484617/
https://jamanetwork.com/journals/jamaophthalmology/fullarticle/2629120
https://slideplayer.com/slide/4483715/
Somogyi, A., Petcherski, A., Beckert, B., Huebecker, M., Priestman, D., Banning, A., . . . Tikkanen, R. (2018). Altered Expression of Ganglioside Metabolizing Enzymes Results in GM3 Ganglioside Accumulation in Cerebellar Cells of a Mouse Model of Juvenile Neuronal Ceroid Lipofuscinosis. International Journal of Molecular Sciences,19(2), 625. doi:10.3390/ijms19020625
Boswell-Casteel, R. C., & Hays, F. A. (2016). Equilibrative nucleoside transporters-A review. Nucleosides, nucleotides & nucleic acids, 36(1), 7-30.
Perland, E., Bagchi, S., Klaesson, A., & Fredriksson, R. (2017). Characteristics of 29 novel atypical solute carriers of major facilitator superfamily type: evolutionary conservation, predicted structure and neuronal co-expression. Open biology, 7(9), 170142.
Schultz, M. L., Tecedor, L., Lysenko, E., Ramachandran, S., Stein, C. S., & Davidson, B. L. (2018). Modulating membrane fluidity corrects Batten disease phenotypes in vitro and in vivo. Neurobiology of disease, 115, 182-193.
Images:
https://www.verywellhealth.com/myotonic-muscular-dystrophy-symptoms-4570998
https://www.ninds.nih.gov/Disorders/Patient-Caregiver-Education/Fact-Sheets/Batten-Disease-Fact-Sheet
https://www.researchgate.net/figure/RNA-seq-data-analysis-of-BMs-AMs-and-TMs-A-Heatmap-of-hierarchical-clustering_fig1_321441357
http://geneontology.org/
https://www.researchgate.net/figure/Effect-of-cisd2-Ppt1-and-cln3-on-external-eye-morphology-Light-microscopic-images-A-K_fig10_261408029
https://www.studyread.com/importance-of-lysosomes/
https://www.sciencedirect.com/science/article/pii/S0167488907000705
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4201705/
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5484617/
https://jamanetwork.com/journals/jamaophthalmology/fullarticle/2629120
https://slideplayer.com/slide/4483715/
This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison. genetics564.weebly.com/