What is transcriptomics? [1]
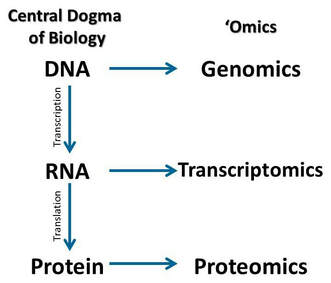
Transcriptomics is the study of the transcriptome; DNA transcripts (DNA copied into RNA). Because an RNA sequence mirrors the sequence of DNA from which it was transcribed, scientists can analyze the entire collection of RNA in a cell and determine which genes are turned on and off in an organism. In humans and other organisms, nearly every cell contains the same genes, but different cells show different patterns of gene expression. These differences are responsible for the many different properties and behaviors of various cells and tissues, both in health and disease. By collecting and comparing transcriptomes of different types of cells, researchers can gain a deeper understanding of what constitutes a specific cell type, how that type of cell normally functions, and how changes in the normal level of gene activity may reflect or contribute to disease. Transcriptomics is often studied via microarrays, qRT- PCR, and RNA sequencing. In the images listed below, GEO profiles were used; a database which stores profiles derived from datasets.
Transcriptomics and CLN3
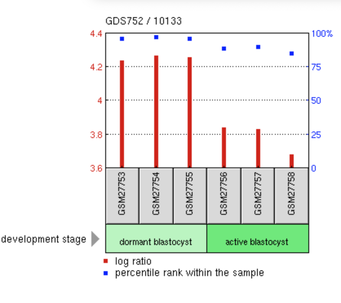
Using GEO Profiles and GEO Datasets, CLN3 expression under different conditions can be analyzed. This could also give insight to what conditions could thwart data.
Conclusion
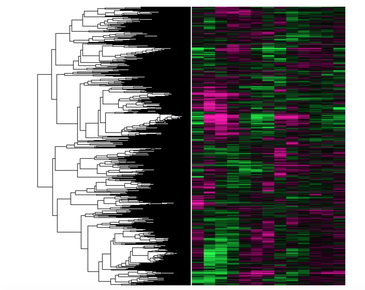
The GEO Profiles and Databases provide insight to which conditions influence CLN3 expression. These are only a few conditions that have been tested, yet could provide insightful knowledge for when the CLN3 gene is over or under expressed in a cell. Also, the heat map comparison has great potential for scientists to infer the function of CLN3 from genes that are expressed in similar manners. For example, if a gene that codes for a transmembrane protein also has high or low expression in the presence of a given condition, scientists can infer that CLN3 is involved with the transcription of that gene.
References:
[1] Transcriptome Fact Sheet. (n.d.). Retrieved from https://www.genome.gov/13014330/transcriptome-fact-sheet/
Header: https://phys.org/news/2016-10-rna-young.html
Figure 1: https://wildlifesnpits.wordpress.com/2015/11/18/transcriptomics-for-conservation/
Figure 2: https://www.ncbi.nlm.nih.gov/geo/tools/profileGraph.cgi?ID=GDS752:10133
Figure 3: https://www.ncbi.nlm.nih.gov/geo/gds/analyze/analyze.cgi?ID=GDS2318
Header: https://phys.org/news/2016-10-rna-young.html
Figure 1: https://wildlifesnpits.wordpress.com/2015/11/18/transcriptomics-for-conservation/
Figure 2: https://www.ncbi.nlm.nih.gov/geo/tools/profileGraph.cgi?ID=GDS752:10133
Figure 3: https://www.ncbi.nlm.nih.gov/geo/gds/analyze/analyze.cgi?ID=GDS2318
This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison